
Svenja Kratz (Recipient of QAGOMA’s 2012 Queensland New Media Scholarship) has recently returned from undertaking a five month residency at the Art and Genomics Centre in Leiden, The Netherlands. Kratz discusses how the award helped furthered her progressive new media practice at the intersection of biology, science and art.
As a child I was always fascinated by biology, particularly organisms that melded with my love of science fiction and horror from zombie snail producing parasites to mutant creatures born with multiple or vestigial organs. These extraordinary organisms challenged my idea of normalcy, embodiment and intelligence, further highlighting evolutionary strategies and myriad interspecies connections. Within my professional practice as an Australian new media artist working across art and science, the idea of the mutant continues to be a point of intense interest. This is partly due to my continued love of the extraordinary, but also because within the context of biotechnology, popular media representations and contemporary science fiction narratives are rife with mutants, from human/pig hybrids proposed for xenotransplantation almost two decades ago, to hybrid creatures of the Splice kind.
These popularist narratives are intense and compelling, but what are the realities of mutant production within contemporary biotechnology, particularly synthetic biology? Are hybrid human/animal creatures really just around the corner? What does it take to make a mutant? As a practitioner, I felt these questions could only be answered through hands-on engagement within the area of genetic engineering and synthetic biology.
Thanks to the Premiere’s 2012 New Media Scholarship from QAG/GOMA, I finally had the opportunity to learn about genetic engineering and mutagenesis during a six-month residency at Gorlaeus Laboratories at Leiden University in The Netherlands from July to December 2013. The residency was organised with support from Robert Zwijnenberg, one of the founding directors of the Art and Genomics Centre and formed part of the large-scale Biosolar Cells research programme. As the title suggests, Biosolar Cells focuses on the potential of solar energy for long term sustainable energy production. While the programme encompasses a variety of research areas including plant research and artificial leaves, I was fortunate enough to be integrated into the Solid State NMR group led by Professor Huub de Groot under the supervision of Professor Wim de Grip and PhD candidate Srividya Ganapathy. The project I worked on aims to increase the absorbance spectrum of light powered protein pumps, which are proteins used by Archaea (single-celled microorganisms) to convert sunlight into chemical energy. Increasing the absorbance spectrum would enable the proteins to use more of the available light spectrum for energy conversion.
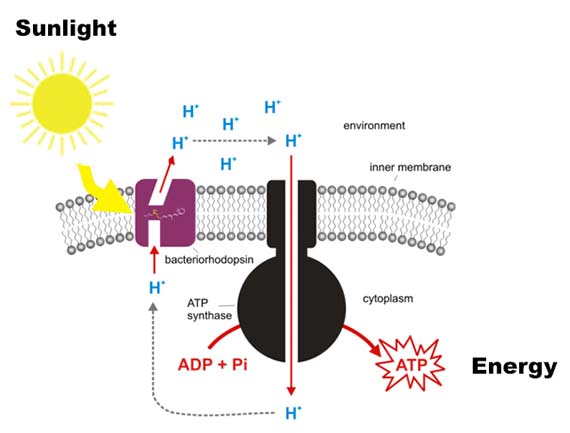
While their research encompasses proteins from a variety of Archaea, my work focused primarily on AR3 – a protein isolated from the organism Halorubrum sodomense. These proteins were incorporated into E.coli bacteria via plasmids (bacterial molecules) – a common genetic engineering method – which enabled them to be grown in large cultures, processed and tested for shifts in absorbance spectrum. The amazing thing is that bacteria with the AR3 protein are able to shift in colour from a standard off-white to stunning purple.
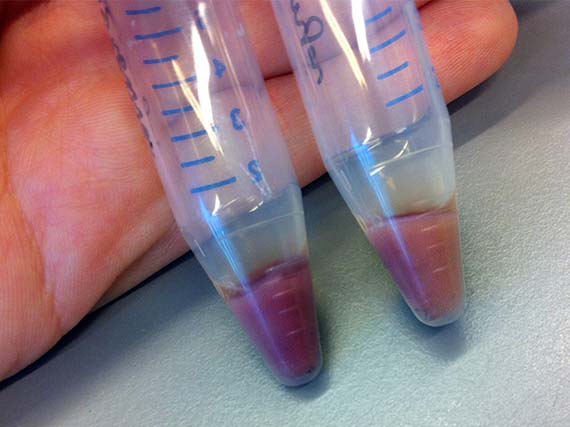
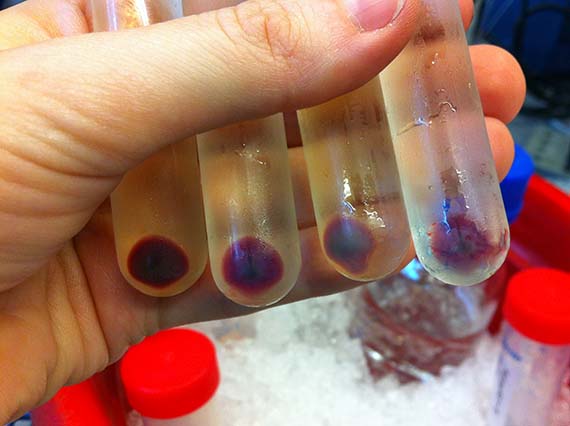
Since, my scientific knowledge was pretty much limited to cell and tissue culture and very basic genetic engineering from my recent PhD studies, the first few months were taken up with training in basic bacterial culture and spectroscopy (a method used to determine shifts in absorbance spectrum), as well as running optimization experiments.
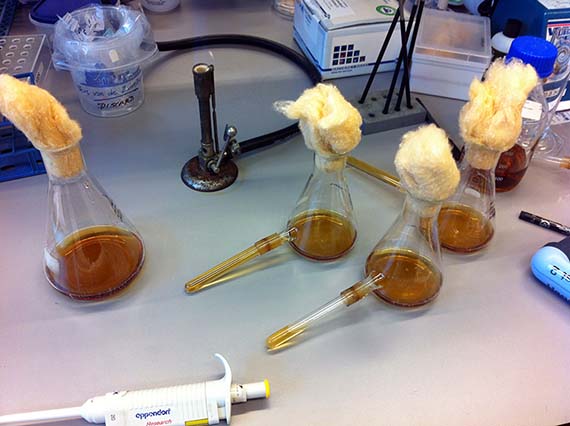
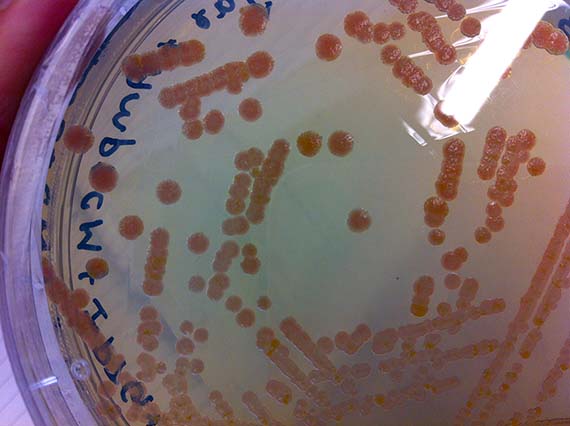

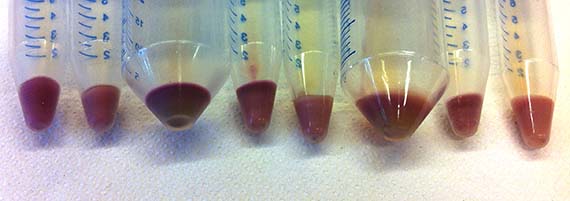
During the initial stages of my residency, I worked hard to impress my lab supervisor and instructor Srividya with my technical competence – largely because she said if I showed enough progress and time permitted, she would show me how to create a mutant. Let me tell you…there is nothing like the promise of mutants to spur on productivity, and with the last months of my residency creeping up, I was finally ready to learn the magical art of site-specific mutagenisis – a common protein engineering process that involves making highly specific changes to a DNA sequence. However, like many laboratory processes, the actual work is far from magical – just many, many long hours pipetting tiny amounts of DNA and chemicals into tiny tubes, amplifying DNA fragments and seemingly endless gels complemented by sleepless nights hoping that the processes were successful.
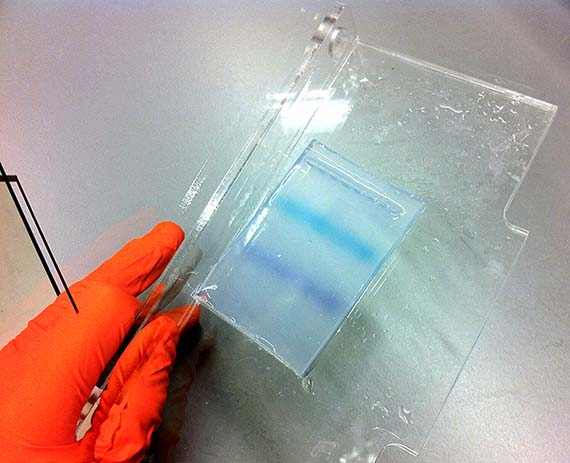

Genetic engineering is definitely not the Frankensteinian melding of parts that I had once imagined, but a somewhat tedious and unglamorous process in which successful outcomes are often invisible to the naked eye and must be determined through other mechanisms such as gel electrophoresis. This process separates macromolecules such as DNA by size and allows the researcher to determine whether the sample contains the right genetic sequence. Indeed, following about a month of labour, the creation of my mutant AR3FS – which included a single amino acid change from phenalynine to serine – was successful. I had succeeded because I had the right bands telling me that the transformation worked!
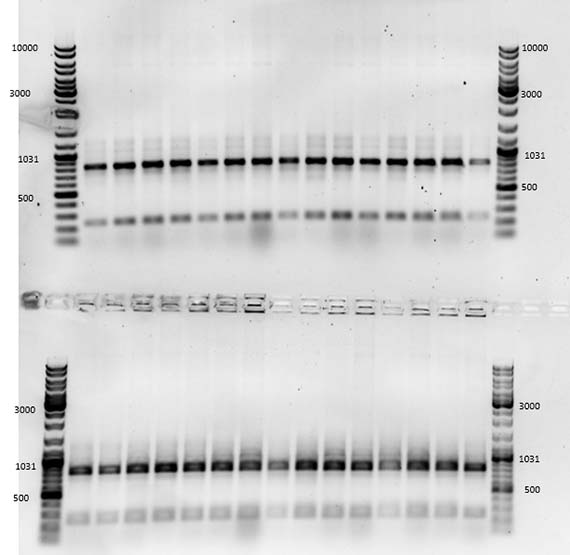
Unfortunately, there were no otherwise discernable changes from the original AR3 – same colour, same absorbance spectrum.
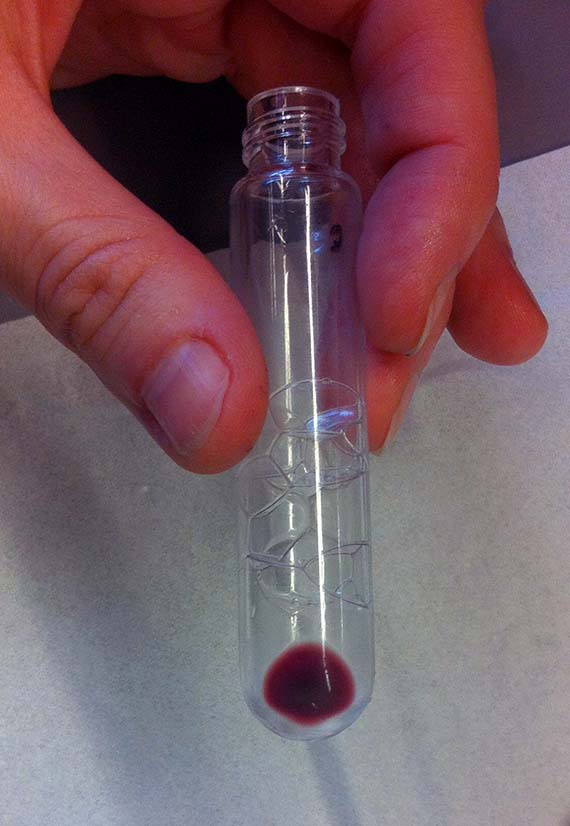
So sadly, my mutant did not live up to my great expectations of significantly advancing the study. I am no science hero, and my mutant is quite frankly a bit of a flop. However, it did show me that many of the real mutants in biotechnology are not large creatures, but often almost imperceptible existing on a micro-scale in the form of bacteria and proteins. It seems that human/animal hybrids are still a long way off and that the real mutants are microscopic (and quite possibly among, and within, us already).
The last remaining weeks of my residency were taken up growing bacterial cultures and isolating the proteins as coloured pigment. Although another tedious process, this enabled me to isolate the protein (no DNA) for safe handling and shipment back to Australia. Indeed, by the close of the residency, I had a whole palette of protein pigments.
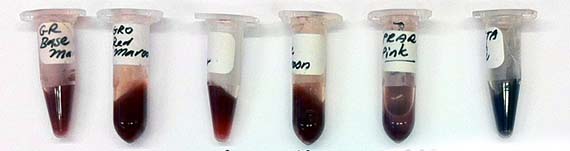
Due to GMO regulations, I was not able to bring any AR3FS samples to Australia, but I prepared a number of frozen stocks prior to my departure. Kept in the -70 freezer at the Gorlaeus Laboratories, they await possible further use and augmentation. I think that is perhaps the most exciting thing about scientific research, the potential of project outcomes, like genetically engineered cells to be used and developed further. I have hope that they may yet contribute something amazing to scientific research.
So…where does the art come in? Well, since finishing the residency, I have completed a series of drawings titled Real and Imaginary Mutants which depict mythological creatures such as the Fiji mermaid and Jackalope, as well as imagined biotech-mutants. These imaginary mutants are coloured with the pigment made from the mutant proteins commenting on the difference between perception and reality of contemporary mutants – within the work, the mutant is the coloured protein and not the representation. These works were exhibited at the SGAR Gallery in June this year along with a series of mixed media works that engaged with the evolution of scientific knowledge and human desire for understanding and control over nature.
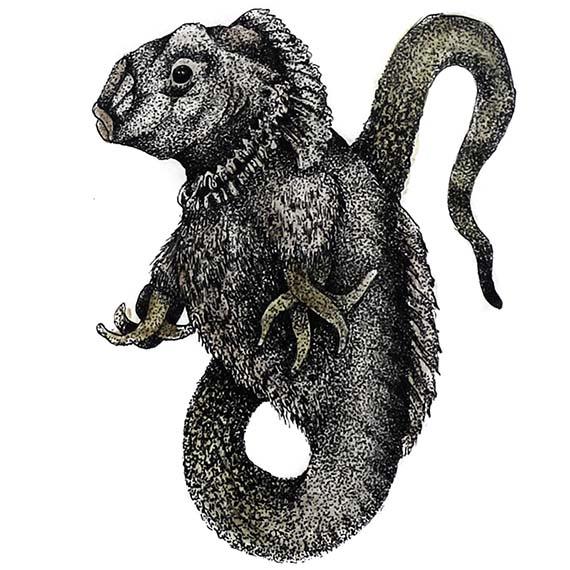

In addition to the drawings and mixed media works, I am currently working with micro-electronics engineer Michael Maggs on holographic and custom projection mapping display systems. To date we have perfected a projection system which will be premiered as part of Experimenta: Recharge this year.
The holographic chamber is next which will connect more directly with the residency featuring a voice responsive video hologram. Holograms and mutants – what more could you want!


Svenja Kratz | Recipient of QAGOMA’s 2012 Queensland New Media Scholarship




